Detecting species using eDNA
Environmental DNA (eDNA) is the collective term for DNA molecules that are released from living or dead organisms in the form of blood, skin, mucous, gametes or faeces and are freely present in the environment. eDNA can be extracted from water or soil samples, and used to detect the presence and relative abundance of target species. Research has shown that eDNA has been successful in detecting species in low numbers (e.g. rare or invasive species). It also has the potential to be a non-invasive alternative to traditional sampling methods.
As part of Reconnect, eDNA samples from rivers are simply collected using sterilised water bottles. The water is filtered on site using a pump. The eDNA collected on the filter is preserved and extracted at a later stage in the laboratory.

Fig. 1 eDNA filtration using a peristaltic pump
To analyse eDNA samples, quantitative Polymerase Chain Reaction (qPCR) techniques are used. Regular PCR, also known as “molecular photocopying” is a common technique used in molecular biology to amplify DNA fragments to obtain millions of copies of DNA. As part of a PCR reaction, primers are used to isolate a particular region of the DNA by binding to a complimentary DNA sequence. This in turn guides the DNA replication process. Primers can be designed so that they only bind to DNA sequences unique to a particular species. Quantitative PCR is similar to regular PCR, however, in addition to using species-specific primers, a fluorescent species-specific probe is added to the reaction. Similar to the primers, the probe will only bind to target sequences of DNA which are unique to a particular species. If DNA from this species is present in the eDNA sample, the probe will fluoresce as the target sequence is amplified. The DNA is quantified against a standard curve, which is generated using known concentrations of reference DNA from a sample of the target species (typically acquired from tissue e.g. muscle, scales or fin clippings). If no DNA from the target species is present in the eDNA sample, the probe will not bind to DNA, and therefore will not fluoresce during the qPCR. The use of primers and probes in qPCR allows for increased specificity and sensitivity of the reaction.
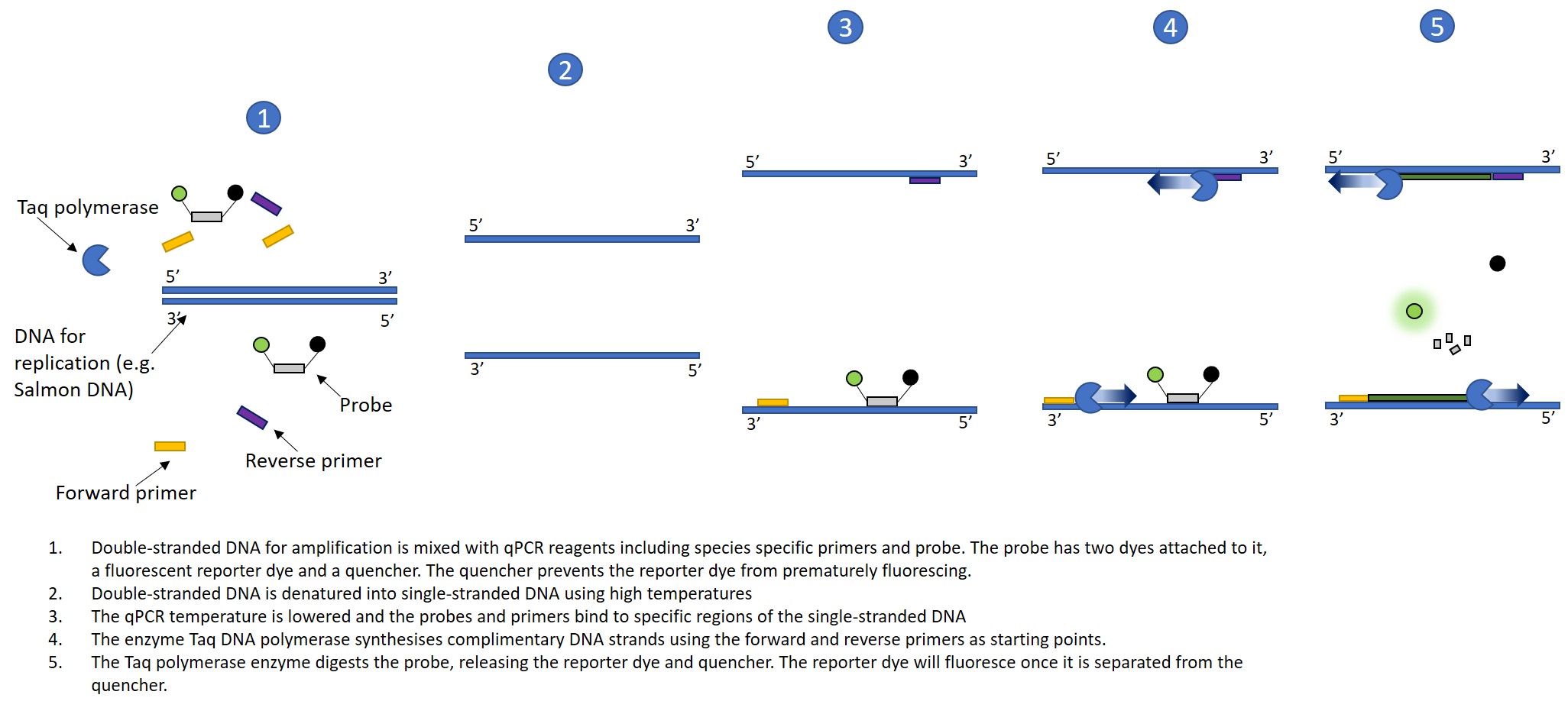
Fig. 2 Diagram showing the steps that occur during a qPCR run. Steps 2-5 are repeated throughout the qPCR run, resulting in exponential amplification of the target DNA sequence. The fluorescence can be measured, giving an estimate of the relative biomass of a particular species in a water sample. As mentioned above, probes and primers will not bind to non-target DNA. In such a case, no amplification would occur, and therefore no fluorescence would be observed. This is why it is necessary to have a quencher attached to the probe as well as a fluorescent reporter dye.
As part of Reconnect, eDNA water samples have been taken upstream and downstream of river obstacles. Primers and probes have been previously developed and deployed for Atlantic salmon (S. salar), brown trout (S. trutta), freshwater pearl mussel (Margaritifera margaritifera), and sea lamprey (Petromyzon marinus). As part of Reconnect, primers for twaite and allis shad (Alosa fallax, and A. alosa), and for the white clawed crayfish (Austropotamobius pallipes) have been developed, and will be deployed in selected rivers.
To learn more about UCD eDNA and population genetics research click the icon below:
