The Good the Bad and the Ugly in Protein Analysis
The aim of proteomics is to completely identify and quantify the entire protein samples that we are interested in. Protein can be isolated from any biological material such as living or conserved tissues, cells, virus particles, or other samples for analytical or preparative purposes like Biopharma molecules and forensic material.
Mass spectrometry (MS) is one of the most popular methods to study peptide and protein characterisation. There are many different methods today to acquire data on a mass spectrometer.
Two different MS strategies are usually applied for the peptide and subsequently protein characterisation:
- Data-dependent acquisition (DDA);
- Data - Independent Acquisition (DIA).
Briefly, the main difference between the two methods are:
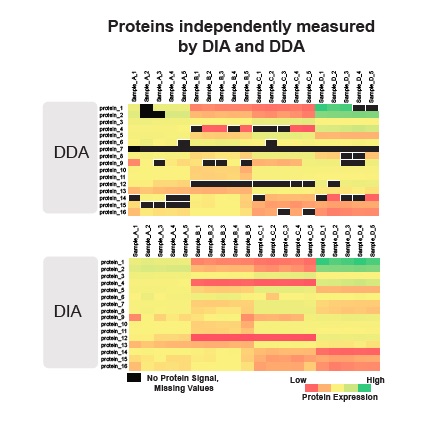
In DDA mode, the mass spectrometer selects the most intense peptide ions in a first stage of tandem mass spectrometry, and then the selected ions are fragmented and analysed in a second stage of tandem mass spectrometry.
In DIA mode, there are no pre-selections. “All the ions are treated equally, no selection is performed.” A predetermined set of wide (often overlapping) scans (isolation windows) are systematically used to send ions and isolate them for fragmentation. Usually, the isolation window coverage is sufficiently wide, all the ions are subjected to analysis.
Many eyes in the proteomics community are now trained on data-independent acquisition (DIA), rather then DDA proteomics mainly for one essential reason:
DIA methods give accurate peptide quantification without being limited to profiling predefined peptides of interest.
Although the DIA concept was introduced a decade ago, interest has been rekindled as several practical DIA implementations have recently been developed. However, further applications to challenging biological questions are needed to showcase the advantages of DIA. Another disadvantage of DIA is the requirement for broader adoption of common and robust data analysis tools, hence DDA methods still mainly used in the Mass Spectrometry Core facilities worldwide.
Why trade my car when it's very reliable? Why switch from DDA to DIA?
Between 2020 and 2021, basically during Covid-time, a positive outcome from working from home gave scientists in the mass spectrometry niche a chance to promote and develop accessible tools for DIA mass spectrometry analysis.
Below are a few papers that summarise the greatest and most interesting platforms developed to support the transition from DDA to DIA currently used in System Biology Ireland. These were uniquely applied by SBI Investigator, Giorgio Oliviero, to investigate phenomena associated with cancer and development with the specific adaptations of cancer with the aging process.
- DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput
- MaxDIA enables library-based and library-free data-independent acquisition proteomics
“Vadim Demichev et all., introduced an easy-to-use integrated software suite, DIA-NN, that exploits deep neural networks and new quantification and signal correction strategies for the processing of data-independent acquisition (DIA) proteomics experiments”
(published in Nature Methods, DOI: https://doi.org/10.1038/s41592-019-0638-x)
“Pavel Sinitcyn et all., introduced,MaxDIA is a software platform for analyzing data-independent acquisition (DIA) proteomics data within the MaxQuant software environment”
(published in Nature Biotechnology, DOI: https://doi.org/10.1038/s41587-021-00968-7)
- IceR improves proteome coverage and data completeness in global and single-cell proteomics
“Mathias Kalxdorf et all., backup the DDA method acquisition which it notoriously suffers from high rates of missing values, thus prohibiting consistent protein quantification across large sample cohorts.
They solve this, with IceR (Ion current extraction Re-quantification), an efficient and user-friendly quantification workflow that combines high identification rates of data-dependent acquisition with low missing value rates similar to data-independent acquisition”
(Published in Nature Communications, DOI: https://doi.org/10.1038/s41467-021-25077-6)
Final thoughts
Some experts believe that the best scenario is the combination of both methods hence DDA and DIA will eventually merge into a single hybrid method, “Data dependent-independent acquisition proteomics," or "DDIA" for short. The results of such integrative methodologies may be imperative for the development of more accurate predictive and prognostic cancer biomarkers and be compatible with clinical diagnostic workflows and will ultimately facilitate the delivery of precision oncology medicine to achieve better patient outcomes.

About the Author
Dr. Giorgio Oliviero is an SBI Investigator who joined the team in 2019.

