Go With Your Gut
I am a 4th year PhD student in SBI and my research for the past four years involved studying colorectal cancer, more commonly known as bowel cancer. The project so far has produced a lot of data and I had the pleasure of presenting this work at the recent 58th IACR Annual Conference in Cork in March of this year, which was a scientific presentation for researchers. In this blog I want to explain my work for a non-specialised audience and to share the relevance of the work I have been doing.
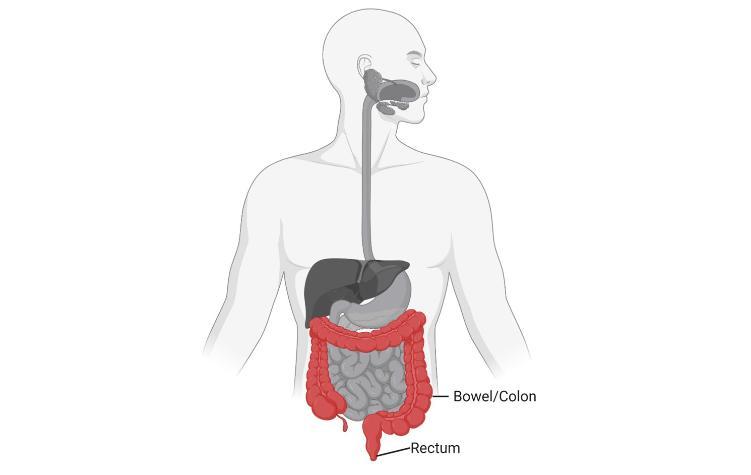
Figure 1: Bowel/Colon and rectum (Colorectal) position in the body (Created with BioRender.com).
Before I get into what I do, I want to give some information about bowel cancer and explain why the work we are doing is important. Bowel cancer is one of the most common forms of cancer in Ireland and worldwide1. It is an extremely difficult cancer to treat due to many patients being diagnosed when the cancer has already spread throughout the body, making it much more difficult to deal with. Many patients are diagnosed at a late stage because symptoms don’t show up early in the disease when it is still limited to the bowel2. This is why screening is important for this type of cancer, as this can sometimes find the disease in the early stages and stop it from progressing3. Although no screening test is 100 percent reliable, it is relatively straightforward and painless, and can be done from the comfort of your own home. Risk of bowel cancer increases with age and in Ireland people aged between 60 and 69 are invited to do these screening tests for free (more information is available on the HSE website).
What causes this cancer? Unfortunately, the number of people developing bowel cancer all over the world is increasing and this is the same in Ireland. We know that there are different factors contributing to this increase but one important one is diet and lifestyle. Diets high in red and processed meats but low in fibre, fruits and vegetables, and vitamin D increase the risk of bowel cancer4. Lifestyle factors include obesity, smoking, and alcohol consumption4. Other factors such as family history, inflammatory bowel disease, polyps, gut microbiota, and socioeconomics also influence the occurrence of this cancer4.
The development of bowel cancer usually follows a sequence that we understand very well. In Figure 2 we can see the development of the cancer from a small benign polyp to a cancer and this process is caused by changes in the DNA of cells that are called mutations. Different mutations are associated with each stage of the disease, which can take years to grow5. My project focuses on KRAS mutations, involved early in the disease (highlighted in the blue box in Figure 2). This mutation is very common in people with bowel cancer and our group has been working on understanding how it causes cancer for a long time.
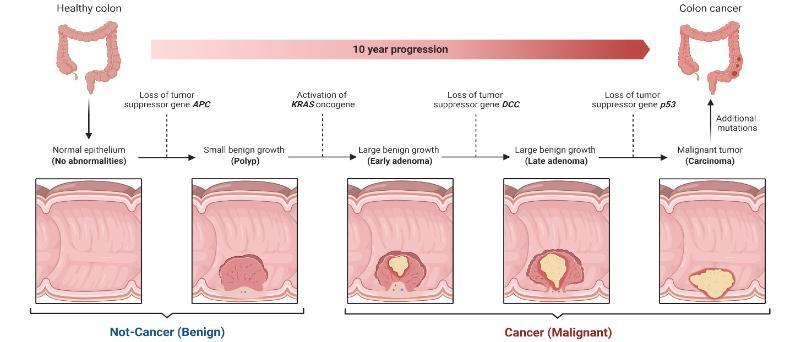
Figure 2: Progression of polyps in the bowel from benign to cancer over time, with mutations at each stage of the disease (Created with BioRender.com)
So what actually is KRAS? KRAS is a gene that is in every cell in our body. This KRAS gene produces a KRAS protein that acts like a switch6 (Figure 3). When the switch is “on”, this leads to cell growth and when the switch is “off”, this leads to no cell growth. Normal healthy cells can find a good balance between turning the switch “on” and “off”. However, mutations can cause the switch to be stuck in the “on” state7, leading to the cell reproducing when it should not, which in the end can result in the build-up of many cells in what we call a tumour.
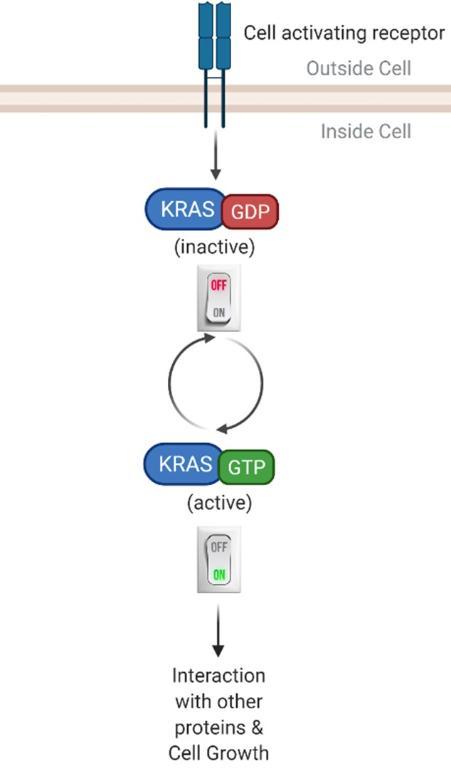
Figure 3: KRAS switch mechanism is balanced in normal healthy cells, but in cancer KRAS mutations are always "on" (Created with BioRender.com).
Until recently, most KRAS mutations were thought to be equal or have the same effect on cells but, based on some previous results in our group, we think that there may be differences among the mutations. Therefore, my project involves looking at some of the different KRAS mutations and how these different KRAS mutations work as switches and what their effect is, if any, in cells and patient tumours.
The first thing I wanted to see is if the different KRAS mutations bind to the same proteins or do they bind to different proteins. In order to look at this, we used a complex technique called mass spectrometry. Basically, this is like fishing with bait. In our case the baits are the different KRAS mutations, and the catch (or fish) are the proteins that bind to them. Using the mass spectrometer, we can see the proteins caught by each bait (KRAS mutation). The different KRAS mutations had some different proteins binding (Figure 4). After quite a bit of analysis, I could see that some proteins bind to one mutant and not to the other. Other proteins bound the two KRAS mutations. We believe that this difference in the “catch” shows that KRAS mutations are different and act differently as switches for certain proteins.
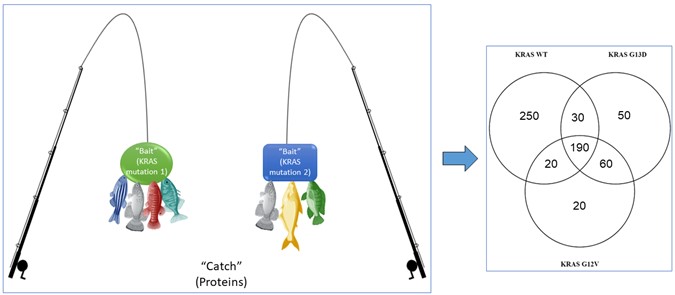
Figure 4: Fishing for proteins that bind or interact with the different KRAS mutations. KRAS mutations are not the same. The Venn diagram shows the results from this analysis with a number of proteins bound with only one KRAS mutation, some proteins bound to both KRAS mutations, and some proteins bound to all three KRAS types (the KRAS wildtype (WT) is our control, as normal healthy cells have this KRAS type) (Created with BioRender.com)
I also looked at how these different KRAS mutations affected patient tumours. This is possible because my project is part of an EU funded project called COLOSSUS where we have access to tumours from biobanks and tumours generously donated by bowel cancer patients in Ireland, France, and Spain. These patients kindly gave us permission to use their tumours for research. COLOSSUS is a multinational project that involves scientists and clinicians all over Europe and you can learn more about it in this video. UCD is one of 13 partners from seven European countries. In a nutshell, the aim of COLOSSUS is to deliver better treatment options through a precision medicine approach for bowel cancer patients, outlined in Figure 5. To do this, partners in COLOSSUS use various techniques to gather information from patient tumours. They then use this information with mathematical and computational models to give better treatment options to these patients.
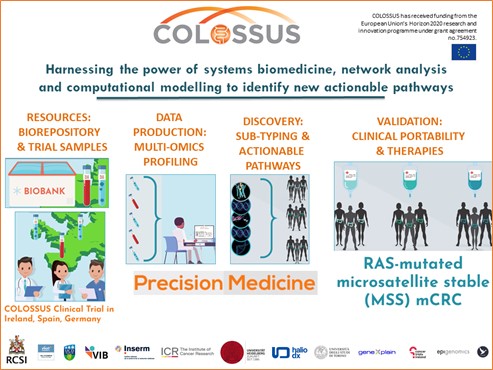

Figure 5: COLOSSUS project overview.
What I have done within COLOSSUS is to use the same technique that I have already mentioned, mass-spectrometry, to see the proteins that are present in the tumours of the different patients. In this case, it is similar to fishing with a net because we want to catch all the proteins in the tumour and then see if there are any differences between the patients (Figure 6). This is because we know that each patient has a unique protein profile, like a fingerprint, and we use this fingerprint to see if we can link proteins and how they are expressed in the tumours with different KRAS mutations. To do this we had to mash up the tumours to get the proteins out of the tissues and into a liquid form. We then cut up the proteins into smaller pieces called peptides, using an enzyme called trypsin. The peptides are what the mass spectrometer measures. From this data we use different computer programmes to analyse the results, as outlined in Figure 6, and group patients accordingly.
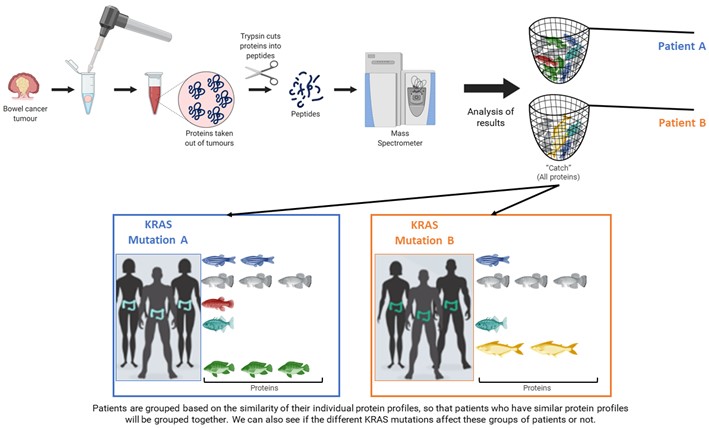
Figure 6: Workflow from patient tumours to protein extraction, mass spectrometer analysis, and the overall aim to use the results to group patients for a more targeted treatment approach to improve patient outcomes (Created with BioRender.com).
Results showed some proteins do associate with certain KRAS mutations, and further work is needed to confirm these results. Interestingly, results from my project so far suggest that KRAS mutations should not be treated the same in patients.
So, why find differences between the KRAS mutations and how does this help patients suffering from cancer? The aim of finding differences among KRAS mutations helps us to find more targeted treatments for patients. The hope is that these targeted treatments are better for patients and improve survival. The more targeted a treatment is, the more efficient it is in stopping the cancer from growing with fewer harmful side effects for patients.

This project has received funding from the European Union's Horizon 2020 research and innovation programme under grant agreement No 754923. The materials presented and views expressed here are the responsibility of the authors(s) only. The EU Commission takes no responsibility for any use made of the information set out.
References:
- Rodriguez-Salas, N. et al. Clinical relevance of colorectal cancer molecular subtypes. Crit. Rev. Oncol. Hematol. 109, 9–19 (2017).
- Dekker, E., Tanis, P. J., Vleugels, J. L. A., Kasi, P. M. & Wallace, M. B. Colorectal cancer. Lancet (London, England) 394, 1467–1480 (2019).
- Burnett-Hartman, A. N., Lee, J. K., Demb, J. & Gupta, S. An Update on the Epidemiology, Molecular Characterization, Diagnosis, and Screening Strategies for Early-Onset Colorectal Cancer. Gastroenterology 160, 1041–1049 (2021).
- Sawicki, T. et al. A Review of Colorectal Cancer in Terms of Epidemiology, Risk Factors, Development, Symptoms and Diagnosis. Cancers (Basel). 13, (2021).
- Brenner, H., Kloor, M. & Pox, C. P. Colorectal cancer. Lancet 383, 1490–1502 (2014).
- Gasper, R. & Wittinghofer, F. The Ras switch in structural and historical perspective. Biol. Chem. 401, 143–163 (2019).
- McCormick, F. K-Ras protein as a drug target. J. Mol. Med. 94, 253–258 (2016).

About the Author:
Aoife Nolan is a PhD student with David Gomez Matallanas's group in SBI and joined the team in 2018.

